Antibiotic resistance is easiest to understand when you can watch it happen. The morbidostat is a feedback-controlled continuous culture device that keeps bacteria near a constant level of growth inhibition by automatically adjusting drug concentration in real time. This turns evolution into a quantitative time series: we can measure trajectories, compare replicates, and connect phenotypes to the mutations that drive them.
Building a morbidostat: an automated continuous-culture device for studying bacterial drug resistance under dynamically sustained drug inhibition
Nature Protocols 8, 555–585 (2013)
Real-Time Observation of Resistance Evolution
Even when outcomes look similar, the underlying genetic routes can be surprisingly structured.
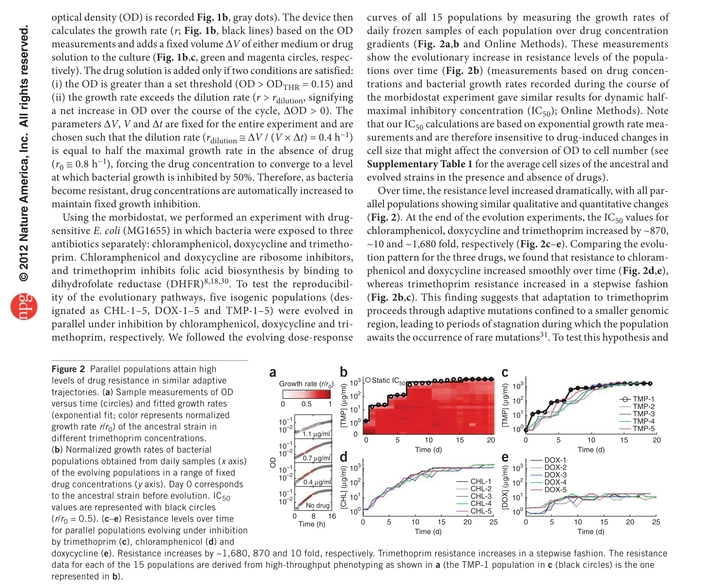
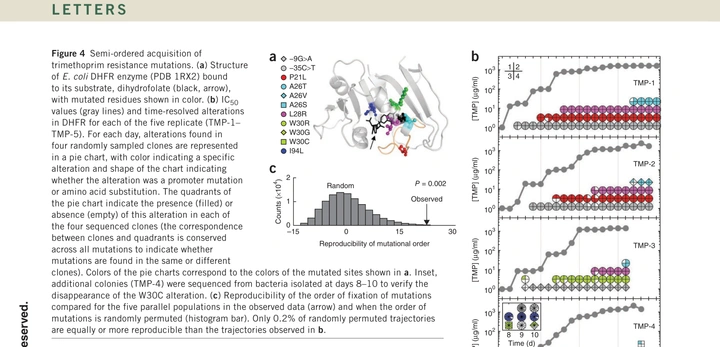
Evolutionary paths to antibiotic resistance under dynamically sustained drug selection
Nature Genetics 44, 101–105 (2012)
The original morbidostat experiments tracked E. coli evolving resistance to multiple antibiotics under dynamically sustained inhibition. Replicate populations repeatedly discovered similar solutions, and sequencing revealed semi-ordered mutational trajectories—a foundation for thinking about resistance as a constrained, measurable process.
Selection Strength Shapes Evolutionary Complexity
Selection strength is a dial: it changes which routes appear, and what they cost.
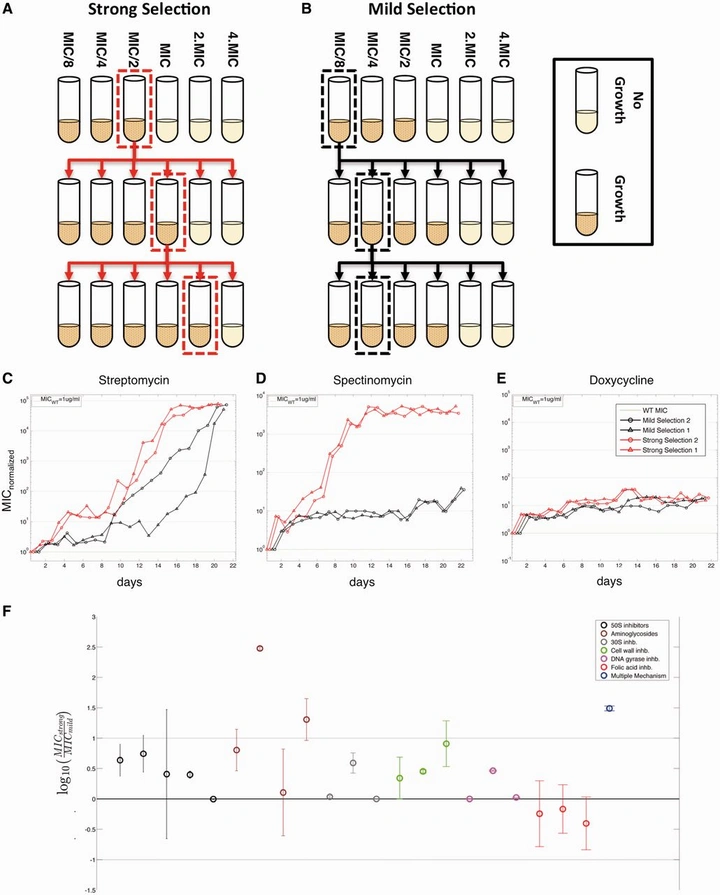
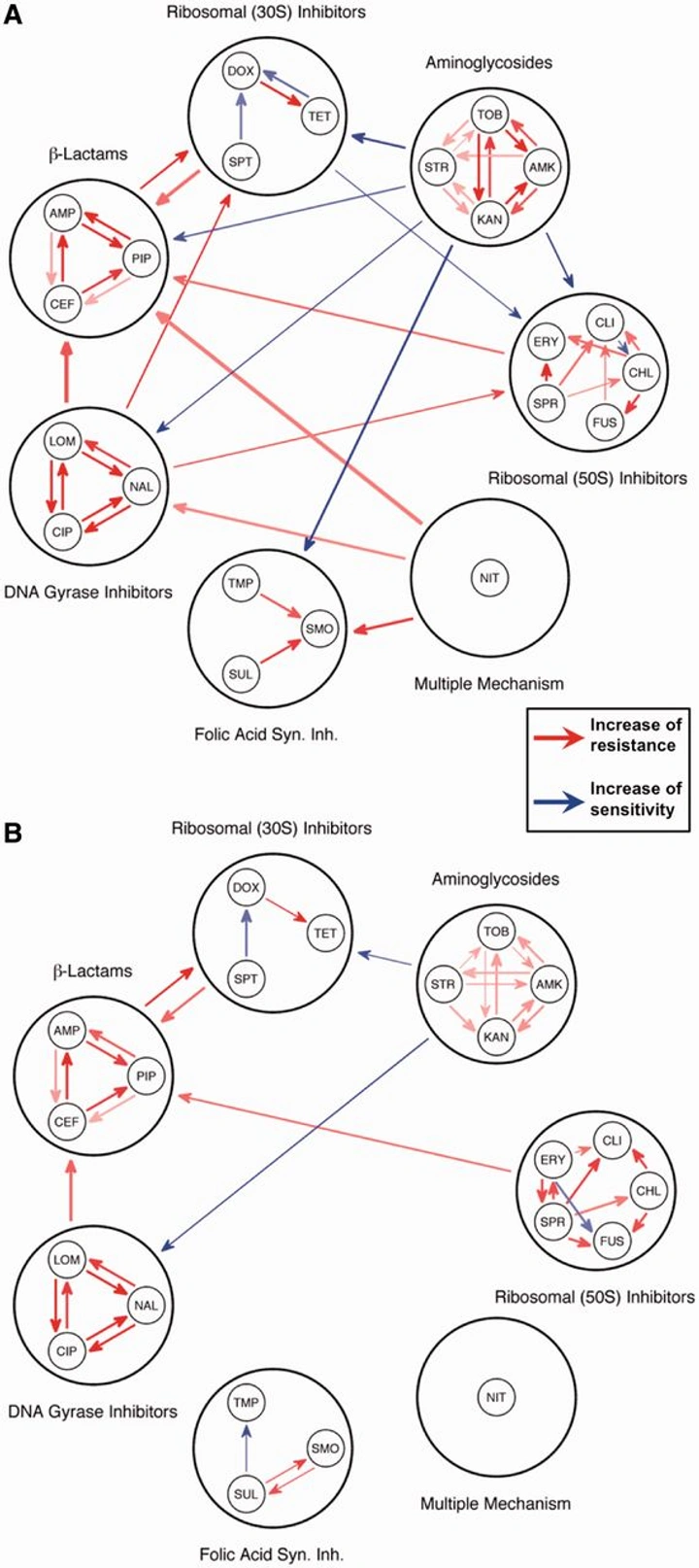
Strength of selection pressure is an important parameter contributing to the complexity of antibiotic resistance evolution
Molecular Biology and Evolution 31(9), 2387–2401 (2014)
By systematically varying selection strength across 88 populations and 22 drugs, this work showed that the intensity of pressure is a key determinant of evolutionary complexity. Similar resistance levels can emerge under strong or mild regimens—but via different genetic routes and with different collateral profiles.