Fast antibiotic susceptibility testing is the missing feedback loop in antibiotic stewardship. We build workflows that turn a blood sample into a quantitative MIC readout in hours—without giving the pathogen a two-day head start: isolate the microbe without waiting for culture amplification, then measure growth with optical sensitivity near the single-cell limit.
Rapid Pathogen Isolation from Blood
Pull the pathogen out of blood in minutes, not days.
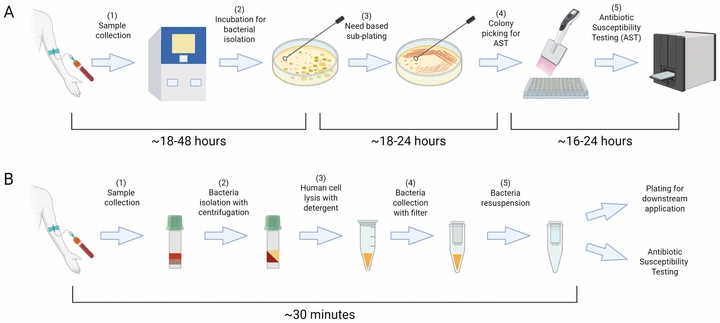
A simple, low-cost, and highly efficient protocol for rapid isolation of pathogenic bacteria from human blood
Frontiers in Microbiology 16:1637776 (2025)
A simple protocol that recovers viable pathogens directly from whole blood in ~30 minutes using standard lab equipment. In validation experiments it achieved >70% recovery efficiency, remained effective at very low bacterial loads, and preserved growth dynamics—so susceptibility testing can start immediately instead of after overnight culture.
Ultrasensitive Optical Detection
Read growth from as few as ~20 bacterial cells.
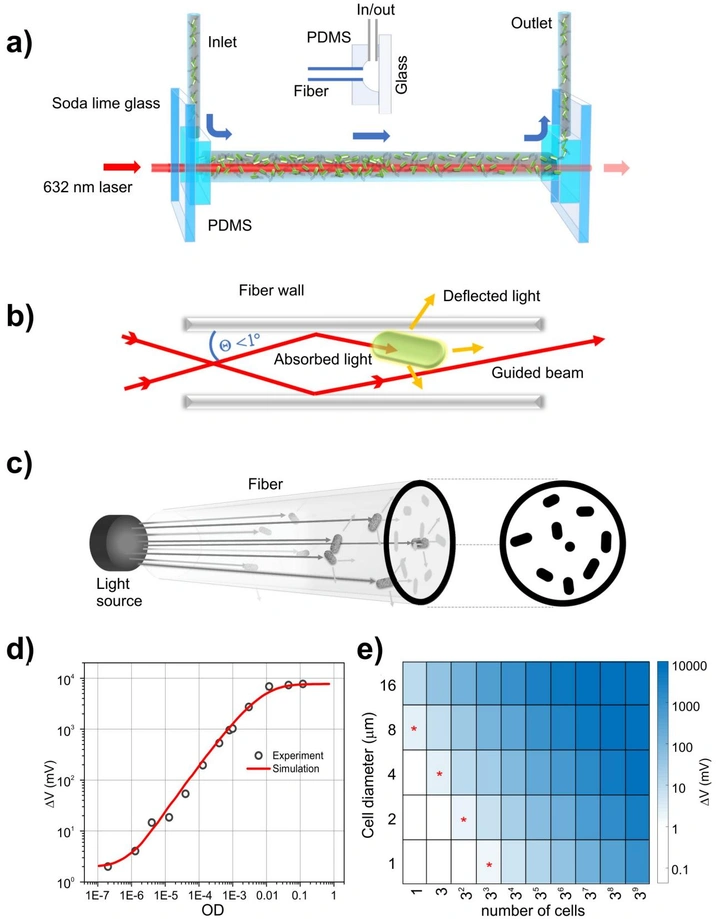
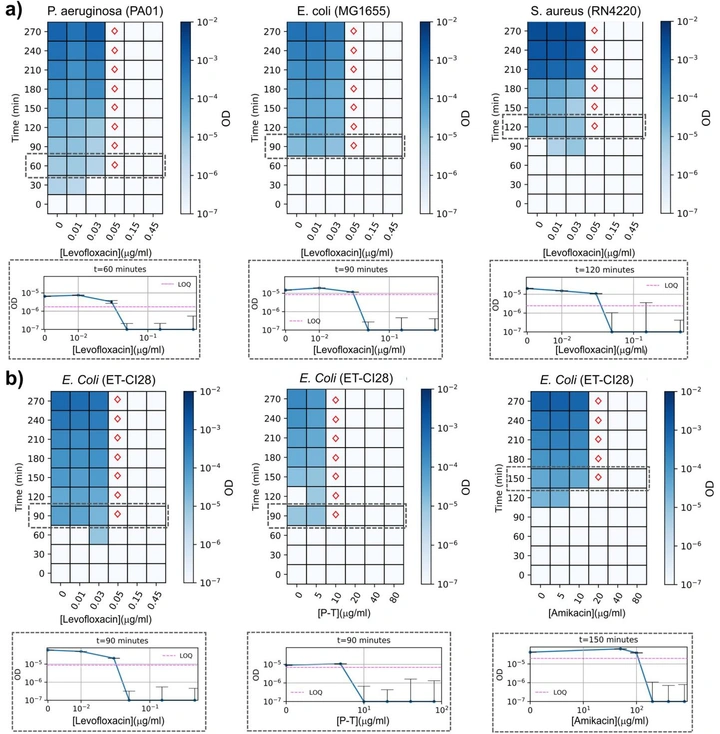
Rapid ultrasensitive detection platform for antimicrobial susceptibility testing
PLOS Biology 17(5): e3000291 (2019)
A label-free optical platform that detects extremely low cell densities (down to ~20 bacterial cells) and enables MIC measurements in 2–4 hours across multiple pathogens and antibiotics. The goal is simple: keep the workflow familiar, but make the readout fast.
From Bench Prototype to Rapid AST
Engineering the optics for robustness, dynamic range, and a clinic-friendly workflow.
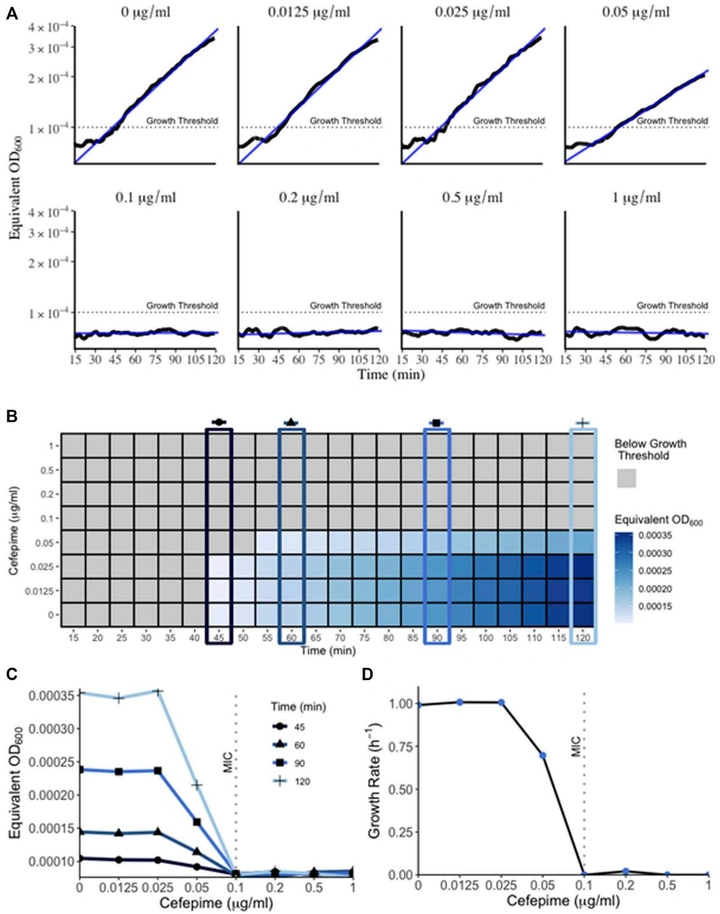
Label-Free Optical Detection of Pathogenic Bacteria and Fungi at Extremely Low Cell Densities for Rapid Antibiotic Susceptibility Testing
Frontiers in Bioengineering and Biotechnology 10:884200 (2022)
Extends RUSD into a more robust apparatus with a wide dynamic window for bacterial and fungal detection and rapid dose-response measurements. In iFAST assays, growth separation can be visible within ~45 minutes and MIC boundaries can be interpreted on the order of an hour—without labels or specialized reagents.