Antibiotic resistance is evolution in a hurry — and it rarely hinges on a single trick. Pathogens can survive by hiding in the host, by taking a few key mutational steps, or by exploring an enormous combinatorial space of enzyme variants. Our goal is to make that search process measurable, so we can predict the paths most likely to appear and design drugs and treatment strategies that close them.
Surviving Without Resistance
Treatment failure can begin with survival—not mutation. We study how susceptible pathogens persist in vivo under clinically grounded drug exposures.
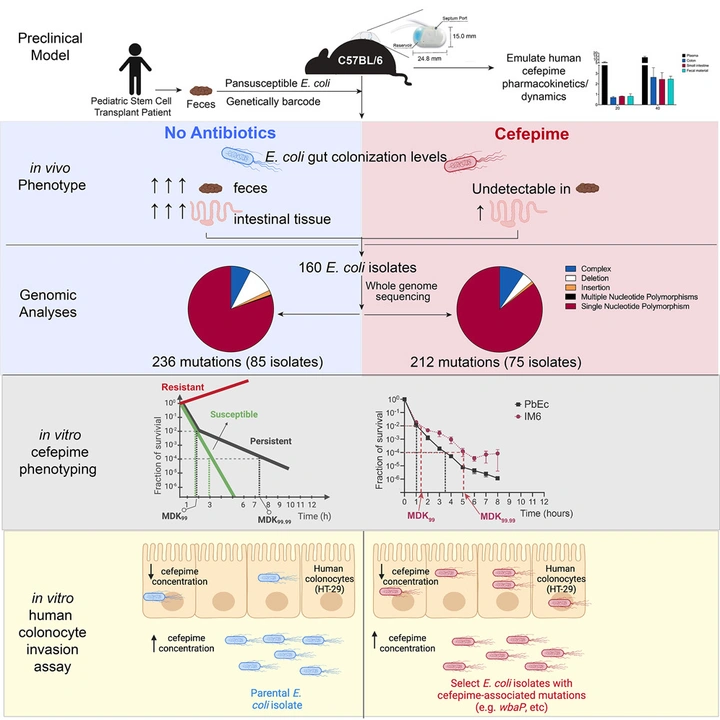
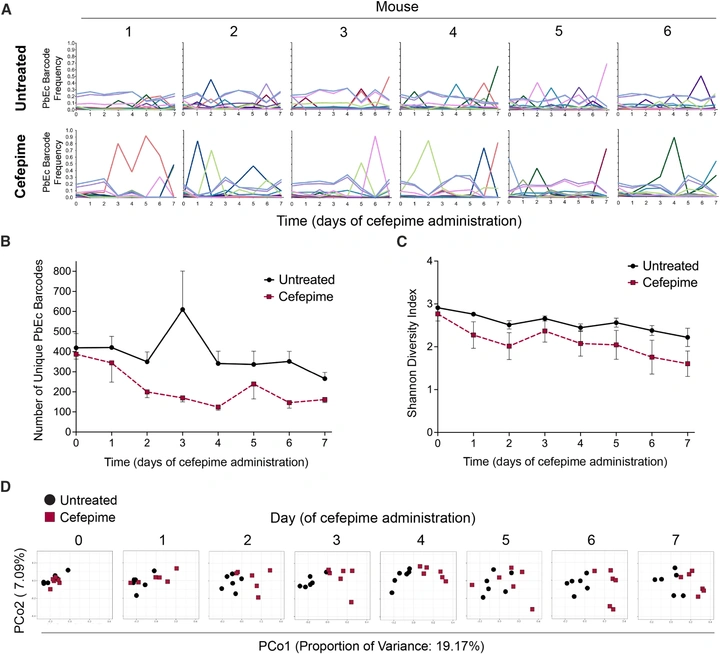
Susceptible bacteria can survive antibiotic treatment in the mammalian gastrointestinal tract without evolving resistance
Cell Host & Microbe (2024)
In a mouse model that emulates human cefepime pharmacokinetics, a pansusceptible clinical E. coli strain disappears from stool yet remains recoverable from intestinal tissue. Survivors do not evolve higher MIC; instead, mutations increase epithelial invasion and/or enable persister-like survival, revealing a non-canonical route to treatment failure.
Steering Evolution with Drug Design
Use molecular mechanism to turn evolution’s favorite shortcut into a dead end.
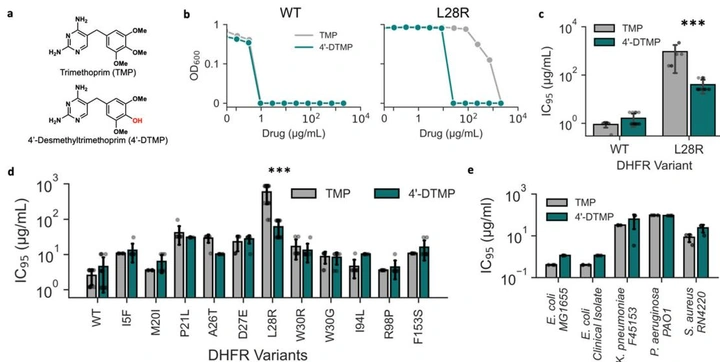
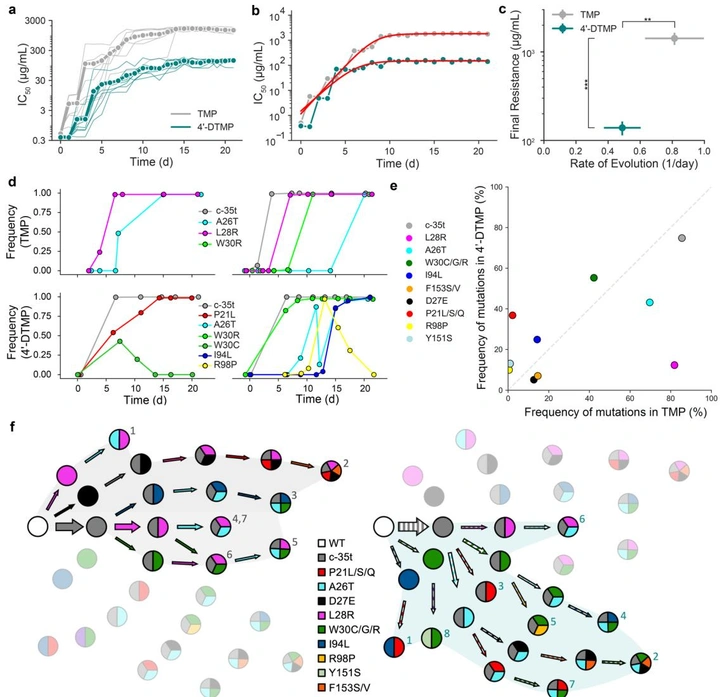
A trimethoprim derivative impedes antibiotic resistance evolution
Nature Communications 12, 3853 (2021)
By dissecting the molecular logic of the canonical DHFR resistance mutation (L28R), we designed 4’-DTMP—an inhibitor that stays potent against L28R. In evolution experiments, 4’-DTMP selects against the usual shortcut and diverts populations to alternative DHFR mutations with catalytic penalties, slowing resistance emergence.
Deep Mutational Scanning
Combinatorially complete fitness landscapes reveal when resistance is predictable—and when it isn’t.
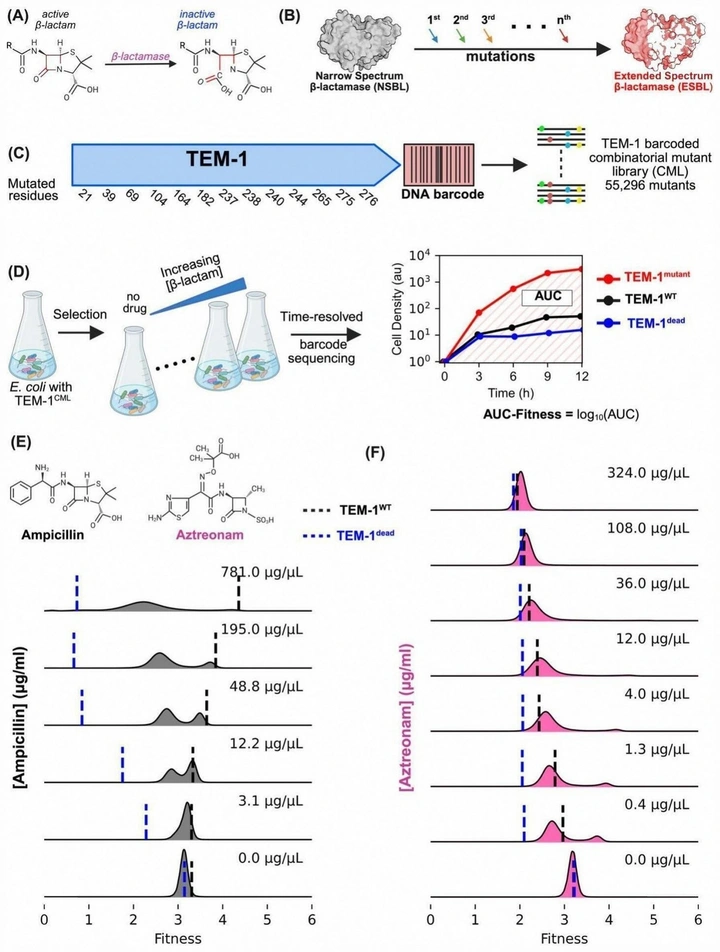
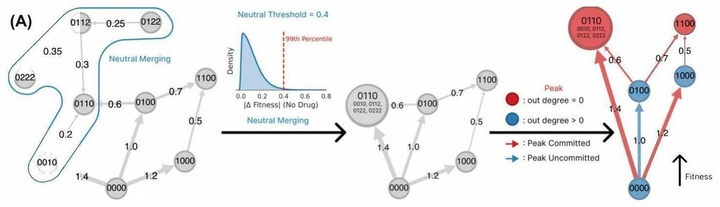
Higher-order epistasis drives evolutionary unpredictability toward novel antibiotic resistance
bioRxiv (2025)
Ilona Gaszek and colleagues measured the largest resistance fitness landscape to date: 55,296 TEM-1 variants with >8 million fitness measurements. Adaptation under a native substrate is comparatively smooth, while selection by a novel drug exposes extensive higher-order epistasis and ruggedness—the root of unpredictable evolutionary outcomes.
Sequential Treatment Dynamics
The same drug can select for different solutions depending on what came before.
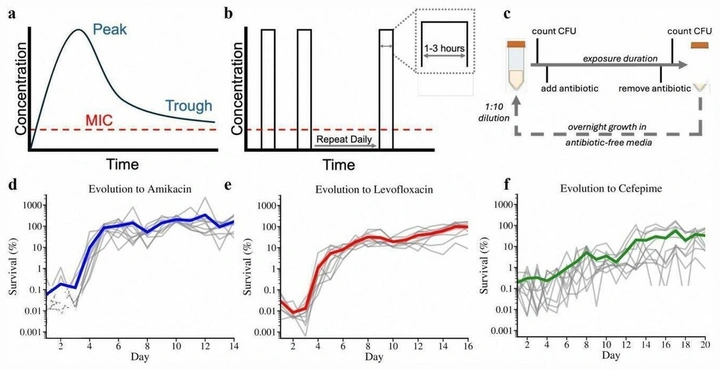
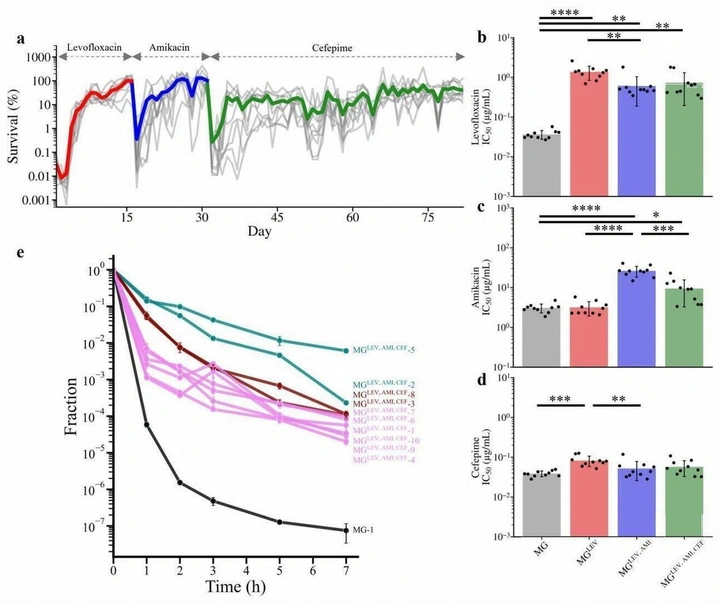
Context-dependent evolutionary dynamics toward bacterial survival during sequential antibiotic treatment
bioRxiv (2025)
Sequential antibiotic use can produce reproducible, history-dependent routes to survival. Our protocol exposes antibiotic-specific solutions (resistance, tolerance, persistence) and shows that treatment order can reshape collateral effects and the mutational paths that remain accessible.